Ring Region –
Relative to Nucleus
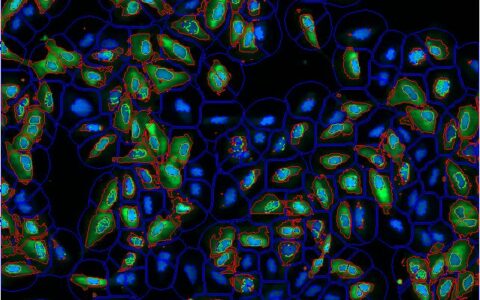
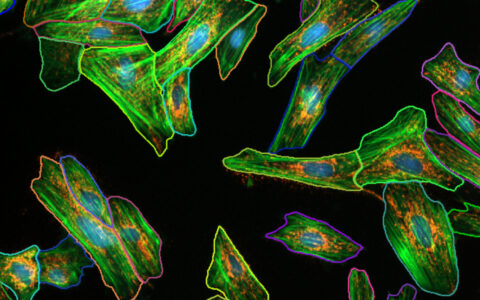
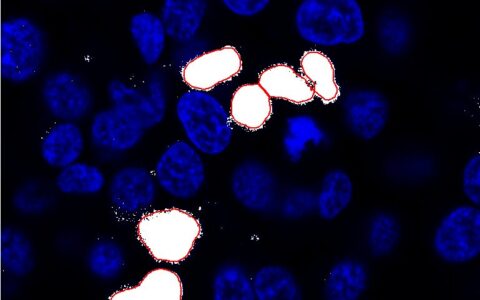
Making cytoplasmic measurements on a per-cell basis is often highly desirable but can be difficult to achieve. This challenge arises when the cytoplasm is not labeled, when cells are densely packed making boundaries unclear, or when the focus is specifically on the perinuclear region of the cytoplasm.
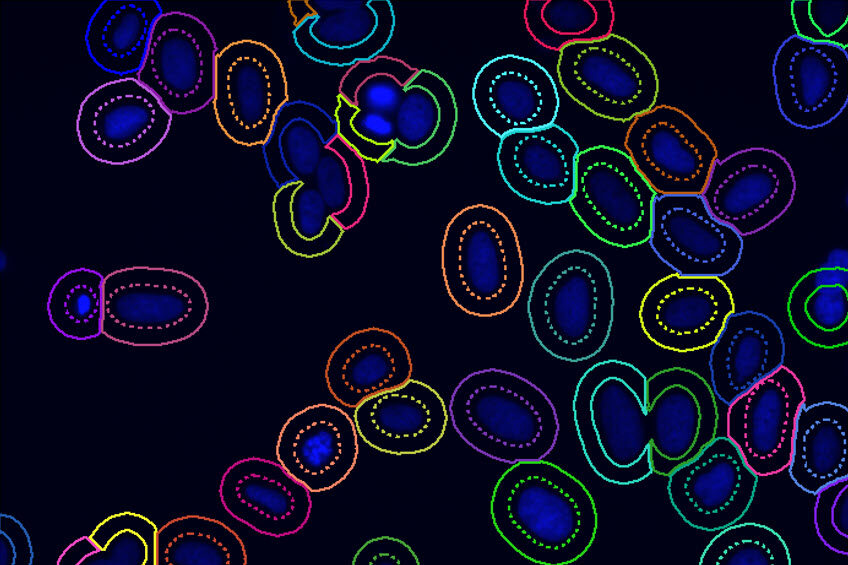
The Image-Pro Ring Region - Relative to Nucleus protocol offers an excellent solution in these cases. It allows you to focus analysis on an annular ring region defined relative to the nucleus, which can be customized based on the position (inside, overlapping, or outside the nucleus), the thickness of the region, and the number of targets you wish to analyze within it. This protocol provides an efficient way to conduct precise cytoplasmic measurements with enhanced accuracy and flexibility.
Techniques: Fluorescence
How it works
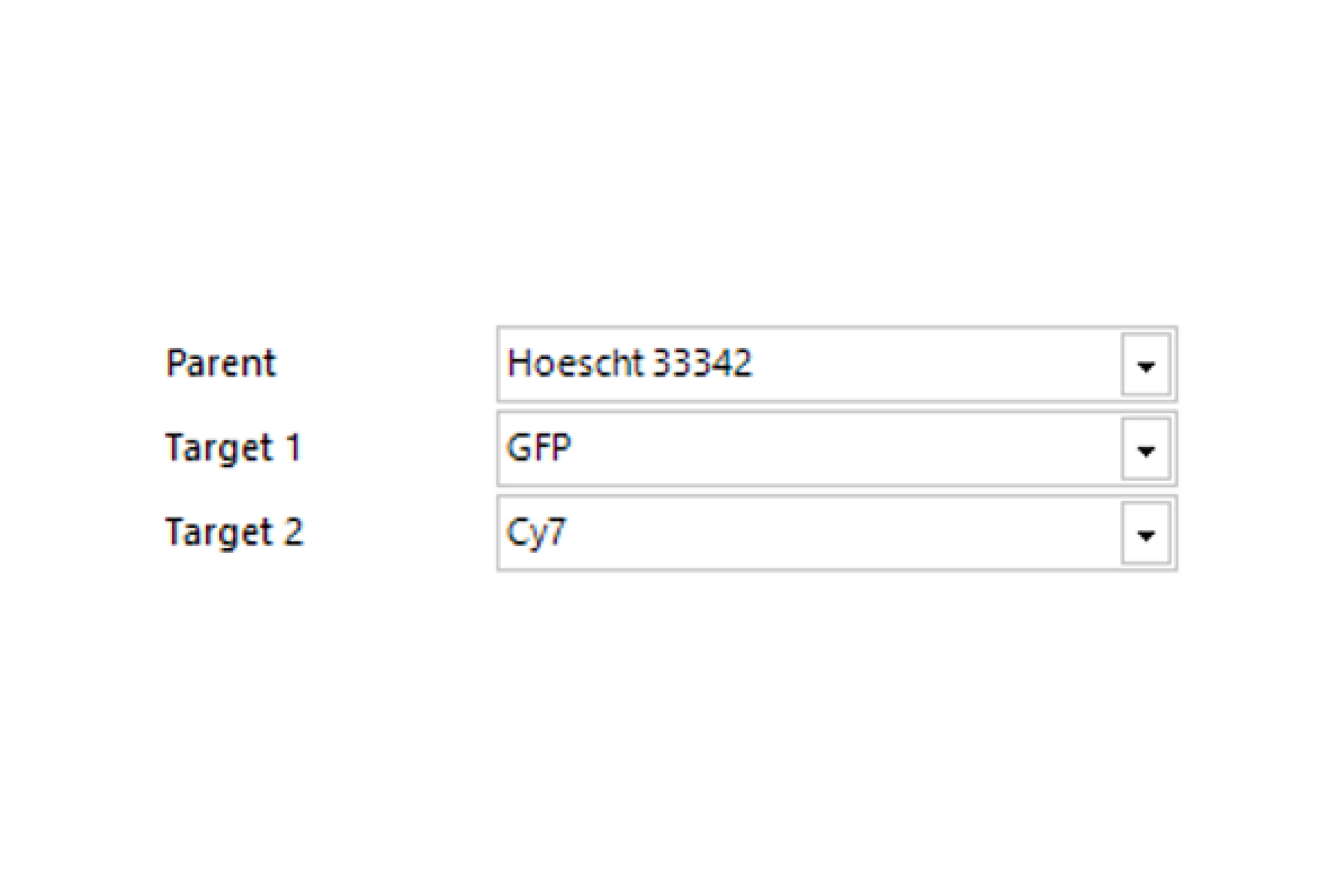

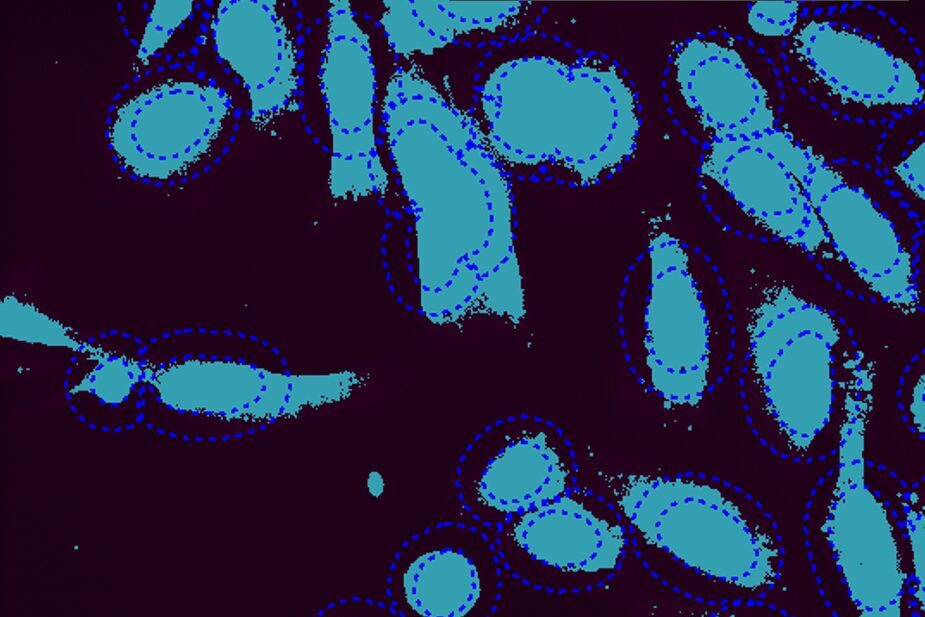
Select Channel
Select the channels that contain labeled nuclei and the target labels or stains.
Find Nuclei
Find nuclei with either a pre-trained deep learning model, machine learning, or threshold segmentation. Define the appearance of the ring region.
Find Targets
Find the first and second targets with either threshold segmentation or machine learning.
Quantitative results
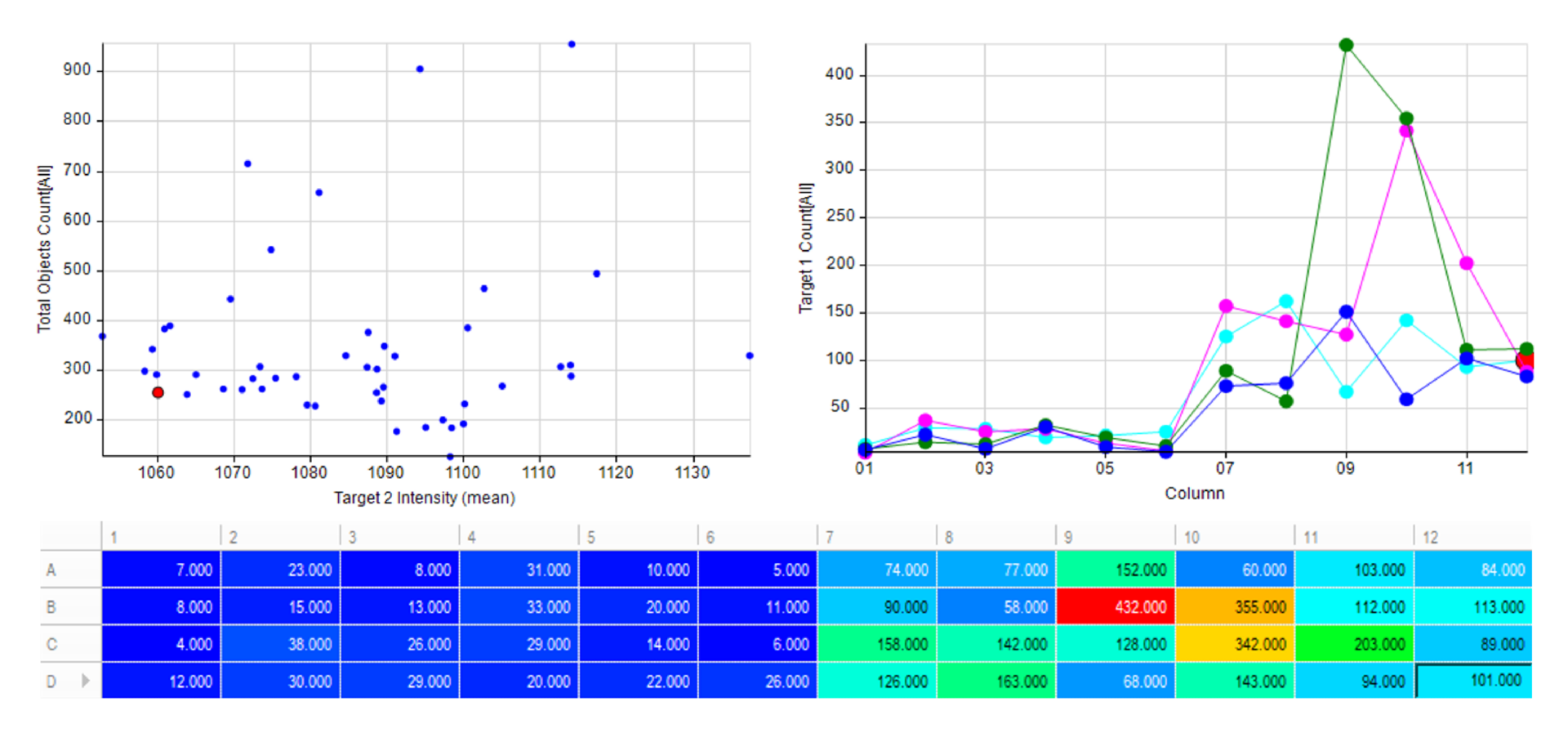
Automatically generate tables, heat maps, charts and even complex bespoke reports.
Measurement parameters supported
- • Target 1 Count
- • Target 2 Count
- • Target 1 Intensity Mean
- • Target 2 Intensity Mean
- • Target 1 Area (Sum)
- • Target 2 Area (Sum)
- • Custom user-defined measurements
Solution requirements
Required Modules
Base
2D Automated Analysis
Cell Biology Protocol Collection
Ring Region Protocol
AI Deep Learning
Life Science Models
Fluorescent Cells Model
Recommended Package